04 May Lesion Detection with Segment Anything Model (SAM)
Segment Anything Model by Meta AI Research is a foundational model for image segmentation. This promptable segmentation model takes points marked on the image, bounding boxes, ambiguous masks or text inputs as an input and gives segmentation labels of objects with associated probability values. SAM also works in a fully automatic mode using a grid to sample all the objects in the image.
SAM consists of three components: (i) image encoder by making use of pretrained Vision Transformer provides image embeddings (ii) prompt encoder works both with points, boxes, masks and text (iii) mask decoder combines image and positional embeddings with a Transformer inspired decoder to create mask(s) with confidence levels.
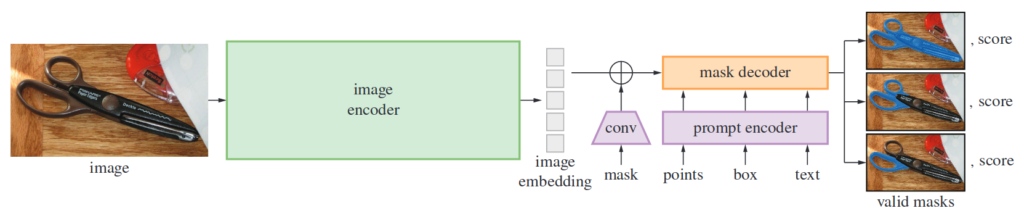
Segment Anything Model (Image taken from Kirillov et al)
SAM is trained on 1.1 billion segmentation images. This massive dataset is created with a blend of model asisted manual annotation, semi and fully automatic segmentation. Final dataset contains more than 1 billion masks segmented from 11 million images.
To examine SAM’s capabilities on medical imaging, we tested SAM on prostate lesion detection. Medical images may contain lower contrast and less variable histogram than RGB images. Also, SAM is not trained on grey level or any type of medical images. This task is particularly challenging for SAM since lesions can be relatively small without any explicit boundaries.
We fine tuned SAM on 1031 MR images with annotated prostate lesions from PI-CAI Challenge dataset [PICAI Challenge]. Prostate lesions are best identified with anatomical and diffusion weighted MR images [PI-RADS 2019]. For this reason, T2 weighted, ADC and High B-value diffusion weighted MR images are concatenated to form three channel volumes. Since SAM works only with two images, from volumetric data axial slices that contain lesions are selected for training and testing. Images are resampled to the same resolution, normalised and resized to make them SAM compatible.
We fine tuned the mask decoder of the model with tight bounding boxes defined around lesions. For finetuning of mask decoder, we followed a similar training recipe as in the original SAM paper. AdamW optimizer (β1=0.9, β2=0.999), with linear learning rate warmup for 250 iterations is used. After warm up, the learning rate is set to 1e-5, weight decay=0.01 is used. Learning rate is decreased by a factor of 10 at 66k and 86k iterations. Batch size was 2. Sum of focal loss (multiplying with factor of 20), dice loss and IOU loss is used for loss function.
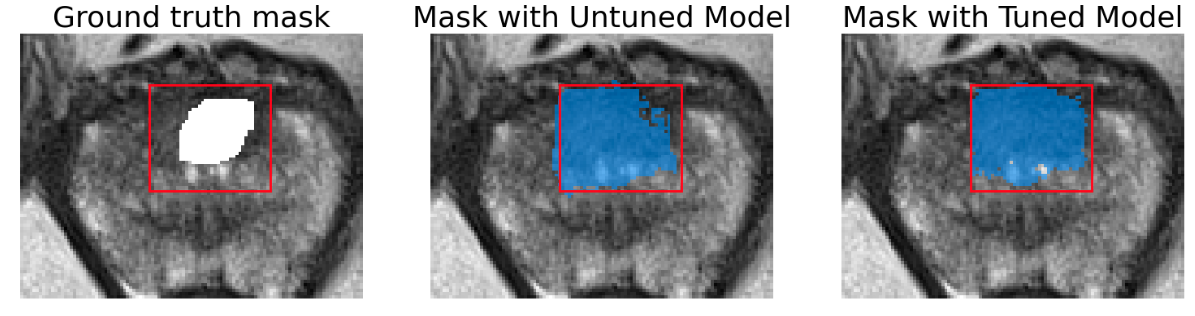
Here we present a few images to demonstrate the performance of SAM on both lesion and prostate segmentation. Results show that tuned model provides only slight improvement and segmented lesions are significantly larger than ground truth.
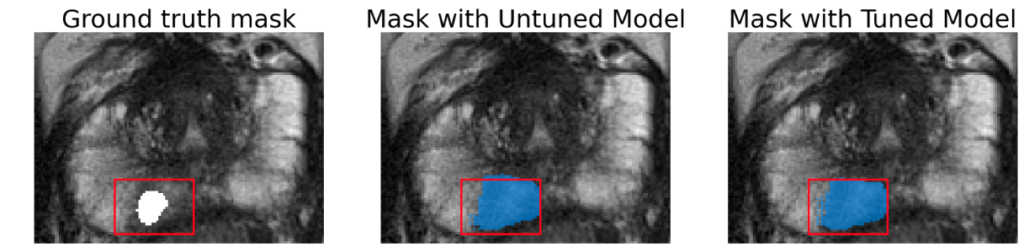
Segmentation area is bigger for both tuned and original model
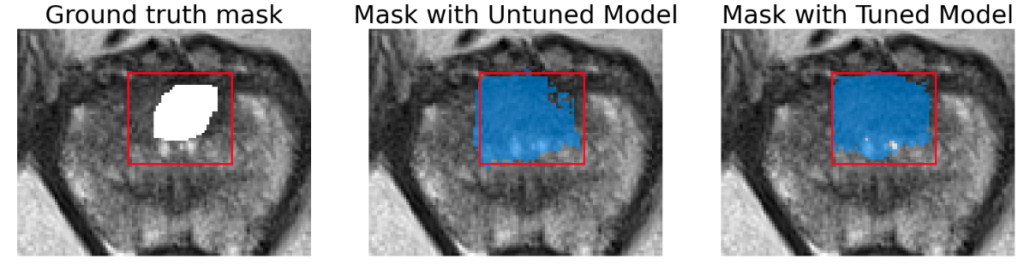
Segmentation area is bigger for both tuned and original model
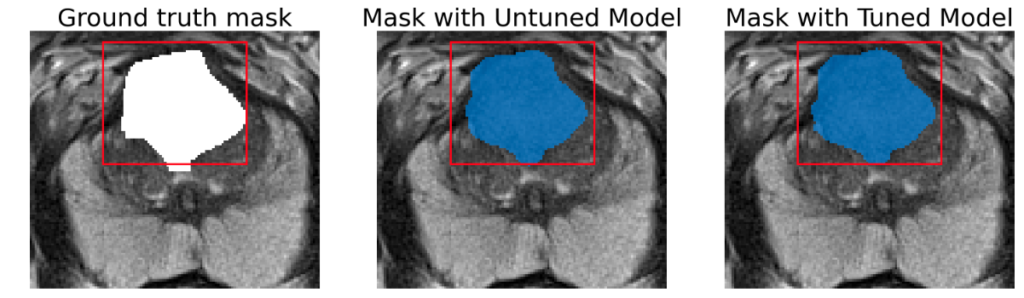
Successful lesion segmentation
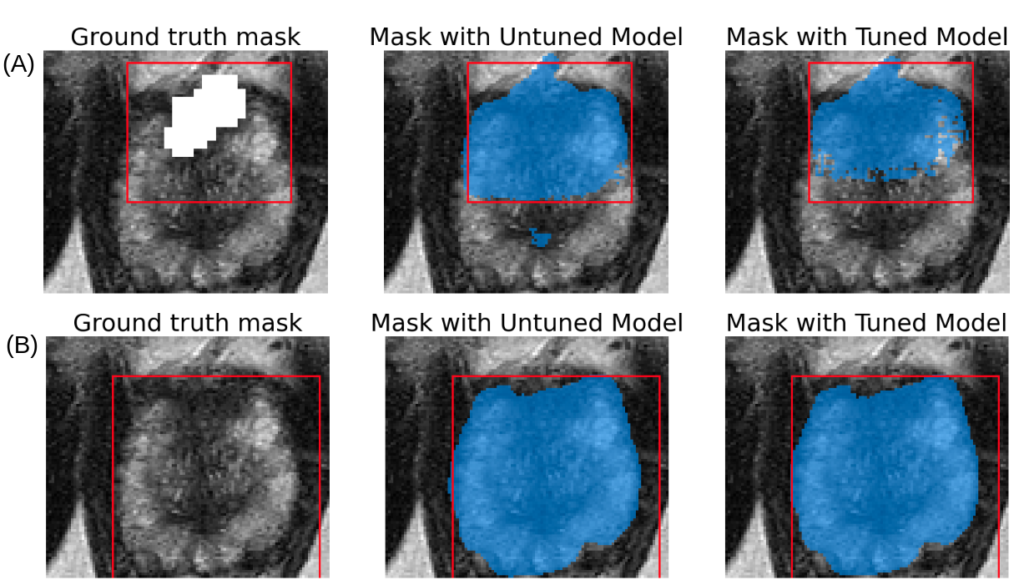
The same image with different bounding boxes are used for testing. In A, the bounding box encapsulates only lesion. In B, the bounding box encapsulates the lesion and the prostate. Though SAM fails for lesion segmentation (A), it performs well on prostate segmentation (B).
Results have shown that although SAM gives remarkable results in RGB image segmentation, its performance on prostate lesion detection needs further improvement. This could be achieved by increasing the diversity of the training dataset with medical images or tailoring the model further.
Reference:
Kirillov, Alexander; Mintun, Eric; Ravi, Nikhila; Mao, Hanzi; Rolland, Chloe; Gustafson, Laura; Xiao, Tete; Whitehead, Spencer; Berg, Alexander C.; Lo, Wan-Yen; Dollár, Piotr; Girshick, Ross, 2023, Segment Anything, http://arxiv.org/abs/2304.02643
PI-RADS 2019 accessed in 3 May 2023 at: https://www.acr.org/-/media/ACR/Files/RADS/PI-RADS/PIRADS-V2-1.pdf
The PI-CAI Challenge: Public Training and Development Dataset https://zenodo.org/record/6624726#.ZFJeppHMKV4